About the course
Course contentsIn this course, you will learn how to analyse single-cell RNA-seq data using the Seurat single-cell tools integrated in the easy-to-use Chipster software. The exercises and course data are based on the Seurat guided analyses "Guided tutorial - 2700 PBMCs" and "Introduction to scRNAseq integration".
This course contains two types of lecture videos: short lectures on each topic by trainers from CSC (ELIXIR-FI), and more in-depth lectures by Paulo Czarnewski (NBIS / ELIXIR-SE), Ahmed Mahfouz (LUMC / ELIXIR-NL) and Jules Gilet (ELIXIR-FR).
You will learn the following topics, and how to perform these steps in the Chipster software:
 perform quality control and filter out low quality cells
perform quality control and filter out low quality cells- normalize gene expression values (with global scaling normalization and SCTransform)
- scale data and remove unwanted sources of variation
- select highly variable genes
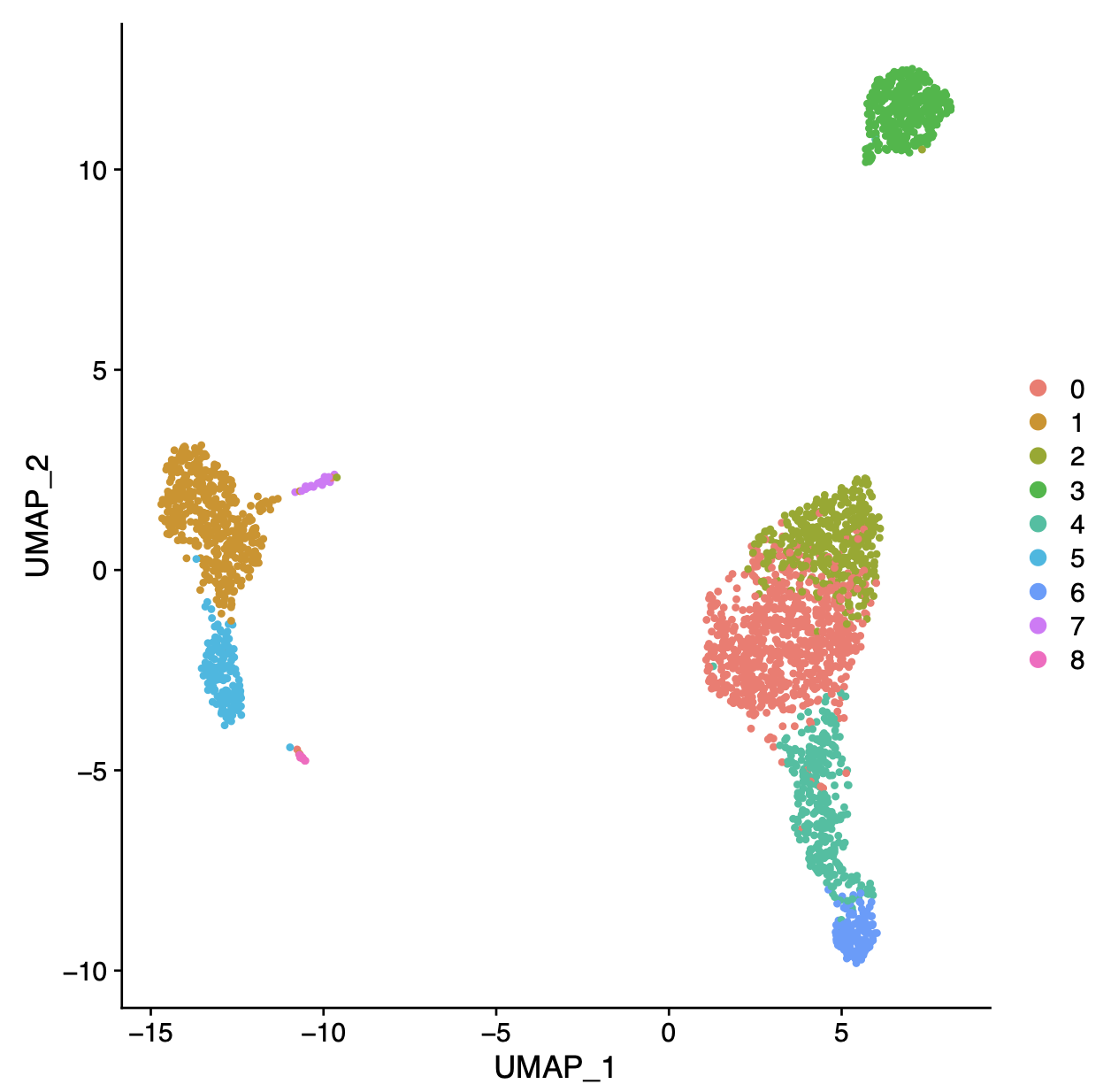
- perform dimensionality reduction (PCA, tSNE, UMAP, CCA)
- cluster cells
- find marker genes for a cluster
- annotate cells and clusters using a reference data
- take a closer look at the Seurat objects
- integrate two samples
- find conserved cluster marker genes for two samples
- find genes which are differentially expressed between two samples in a cell type specific manner
- visualize genes with cell type specific responses in two samples
"It is so nice to be able to do the whole workflow in Chipster, compared to the old model, where I had to transfer the tsv file to R-studio and run Seurat there. -- I learned how to use the Seurat tools in Chipster and what all the steps really mean. I learned to check the results after every step to adjust the next steps parameters and to test different PCA plotting tools. I also learned how to find different genes in the clusters and how to visualize them. I never got this far using the R-pipeline. " -Pinja, course participant & PhD student from University of Helsinki
Learning objectives
After this course you should be able to:
- use the Seurat tools available in Chipster to undertake basic analysis of single-cell RNA-seq data
- name and discuss the different steps of single-cell RNA-seq data analysis
- understand the advantages and limitations of single-cell RNA-seq data analysis in general and in Chipster
Keywords: Chipster, Seurat, single-cell sequencing, RNA-seq, clustering, aligning cells, cluster markers
Links to material
The relevant material is linked in each course section. Here are some quick links:
- Course slides, exercises and other material (available also after the course)
- How to access Chipster
- Seurat tools guided analyses
- Article about sample integration in Seurat
Practicalities
Each section of this course contains lecture videos, hands-on exercises and questions/tasks. The tasks can be used to confirm that you have reached the learning goals. You can use the Q&A Forum below to ask questions regarding the course topics or the exercises. Once you have finished all the tasks, you can download a course certificate with a unique course identifier. You can follow your progress with the progress bar on the right. The estimated time to complete the course is 2-3 days. In the certificate we recommend granting 1 credit (ECTS) for the course.
Help
In practical matters, please contact event-support (at) csc.fi, and in content related questions, chipster (at) csc.fi. You can also join the Weekly CSC research user meetings in Zoom to discuss course matters and get help with the exercises.